plant disease detection
تفاصيل العمل
plant disease detection application built using PyTorch, multi-head attention, MobileNet, ShuffleNet, and a Tkinter GUI for image input. Here's a breakdown of what each part does:
Model Components
MobileNet and ShuffleNet
Pretrained architectures are partially customized by replacing the last layers with convolution or identity layers.
Used as feature extractors within the custom network.
SimpleResidualBlock
A basic residual block using two convolutional layers with skip connection.
ConvBlock
A reusable block of Conv → BatchNorm → ReLU, optionally followed by MaxPool.
ImageClassificationBase
Base class that includes training and validation steps, accuracy calculation, and logging.
? Main Model – ResNet9
This model combines:
A standard CNN backbone (ConvBlock + residual connections).
Feature fusion from:
A customized ShuffleNet (sh)
A customized MobileNet (mobile)
The CNN stream
Three multi-head attention layers, one for each stream, helping the model to focus on relevant spatial features.
The outputs are flattened and concatenated before going through a Linear layer for final classification (38 plant disease classes).
? Model Loading and Inference
Loads a trained model checkpoint: multi (1).pth.
Uses torch.load to restore the model weights.
Applies softmax and gets the class prediction based on the maximum value.
️ GUI with Tkinter
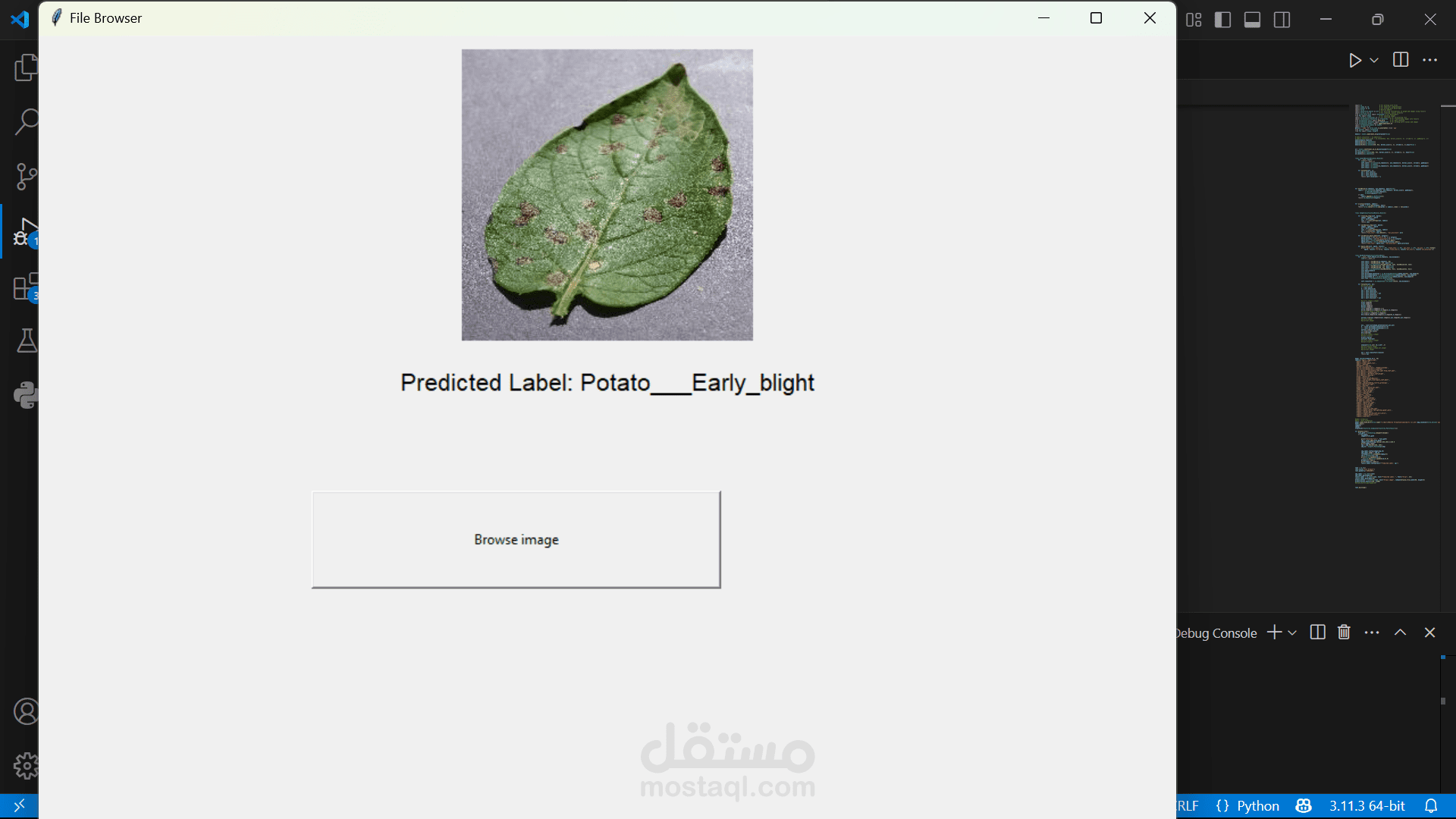
A simple UI to browse an image from the file system.
When an image is selected:
It is resized and transformed into a tensor.
It is passed to the model for prediction.
The predicted label is displayed on the GUI.
️ Classes
labels contains the names of 38 possible plant diseases or healthy categories.
Highlights
Multi-model feature fusion using MobileNet and ShuffleNet.
Multi-head attention for learning richer feature representations.
GUI interface allows practical usage by non-technical users.
Entirely custom model: no pretrained classifier head used.